Maping of Beef Herds in California
- Inquiry commodity
- Open up Access
- Published:
Phylogenomic assay of Mycoplasma bovis from Belgian veal, dairy and beefiness herds
Veterinary Inquiry book 51, Article number:121 (2020) Cite this article
Abstract
M. bovis is one of the leading causes of respiratory affliction and antimicrobial employ in cattle. The pathogen is widespread in different cattle industries worldwide, but highest prevalence is institute in the veal manufacture. Noesis on M. bovis strain distribution over the dairy, beefiness and veal industries is crucial for the design of effective command and prevention programs, but currently undocumented. Therefore, the present report evaluated the molecular epidemiology and genetic relatedness of G. bovis isolates obtained from Belgian beef, dairy and veal farms, and how these relate to M. bovis strains obtained worldwide. Full genomes of one hundred Belgian M. bovis isolates collected over a 5-year period (2014–2019), obtained from 27 dairy, 38 beef and 29 veal farms, were sequenced by long-read nanopore sequencing. Consensus sequences were used to generate a phylogenetic tree in guild to associate genetic clusters with cattle sector, geographical area and year of isolation. The phylogenetic analysis of the Belgian M. bovis isolates resulted in 5 major clusters and i outlier. No sector-specific M. bovis clustering was identified. On a globe scale, Belgian isolates clustered with Israeli, European and American strains. Different G. bovis clusters circulated for at to the lowest degree 1.5 consecutive years throughout the land, affecting all observed industries. Therefore, the loftier prevalence in the veal manufacture is more likely the event of frequent purchase from the dairy and beef industry, than that a reservoir of veal specific strains on farm would be. These results emphasize the importance of biosecurity in G. bovis control and prevention.
Introduction
Mycoplasma bovis (M. bovis) causes mostly pneumonia, arthritis, otitis in calves and mastitis in adult cattle [one, 2] resulting in high antimicrobial use (AMU) and enormous economic losses in cattle farming sectors worldwide [two,3,4]. In Kingdom of belgium, 100% of the veal farms are seropositive for M. bovis [5, half-dozen], whereas M. bovis is involved in 33% of acute pneumonia outbreaks in beef and dairy farms [7]. Treatment of G. bovis is ofttimes unsatisfactory, probably due to a combination of intrinsic and acquired antimicrobial resistance, immuno-evasive backdrop of the pathogen and failure of the animal to generate an constructive immune response [8, 9]. Together with the absence of an effective commercially available vaccine, the control of G. bovis is particularly challenging.
A gimmicky fear is that the veal sector, currently combining a high AMU and a farm level M. bovis prevalence of 100%, is a reservoir for multi-resistant sector-specific M. bovis strains [6, ten, xi]. Currently, there is insufficient knowledge almost the epidemiology of circulating M. bovis strains to answer this question. Several epidemiological studies observed clonal emergence and identified ascendant lineages of the M. bovis bacterium, based on antimicrobial resistance patterns and dissimilar strain typing methods [12,13,14,15]. In the past, unlike approaches were used to subtype 1000. bovis strains, including random amplification of polymorphic Dna (RAPD), arbitrarily primed PCR (AP-PCR), amplification fragment length polymorphism (AFLP), pulsed-field gel electrophoresis (PFGE), multiple-locus variable-number tandem echo (MLVA), and multi-locus sequence typing (MLST) [13, xiv, 16,17,18]. Unfortunately, results from these typing methods are hard to compare and only focus on a small fraction of the genomic information, resulting in a limited insight in the genetics and incongruence among studies [17, xix, xx]. Therefore, whole genome sequencing (WGS) could be a smashing opportunity, because its highly discriminative capacity and reproducibility compared to older typing methods [21, 22].
Several studies already investigated whether specific M. bovis strains were associated with affected organs, such equally udder, respiratory tract or joints [xiv, 23, 24], geographical location [23, 24] or wellness status [23, 25]. Simply one study determined epidemiology based on AP-PCR in three farms from 3 unlike husbandry conditions (dairy dogie ranch, feedlot and closed beef herd), presuming that management factors could influence the distribution of Chiliad. bovis [xvi]. These husbandry weather condition are not comparable with the three master sectors in Europe. In Europe, a lot of short-altitude movements of cattle between farms is seen, and the veal industry is an important side market of the dairy and beefiness industry [26, 27]. It is currently not clear whether sector-specific Thousand. bovis strains are present and what their genetic relation is to previously sequenced Thousand. bovis isolates. Therefore, the present study first evaluated the molecular epidemiology and genetic relatedness of Yard. bovis isolates obtained from Belgian beef, dairy and veal farms. Furthermore, it studied the relationship of these isolates to G. bovis strains from other countries.
Materials and methods
Mycoplasma bovis collection and identification
One hundred M. bovis isolates were obtained from 94 Belgian farms (27 dairy, 38 beef and 29 veal) over a v-year period (2014–2019). All isolates were obtained from diagnostic samples collected by field veterinarians from clinical cases, in compliance with EU legislation on ethics in animal experimentation [2010/63/EU]. Isolates were nerveless in 2014 (north = i), 2016 (11), 2017 (63), 2018 (19) and 2019 (6), originating from the provinces East-Flemish region (n = x), Due west-Flanders (25), Antwerp (38), Limburg (half dozen), Flemish Brabant (10), Heynowes (half dozen), Namen (2) and Liege (1). The origin of two isolates was unclear. The samples were retrieved from the respiratory tract (89), middle ear (3), milk (4), joint (ii) and other fluids (3) of calves and adult cattle, as shown in Boosted file 1. The samples were cultured on selective indicative agar [28], and identification was verified with MALDI-TOF MS (score value ≥ 1.7), every bit described earlier [29] and Kraken2 analysis. Isolates had a passage history of maximum 3–5 times and all isolates were stored at maximum − 20 °C until farther analysis.
Preparation and Deoxyribonucleic acid extraction
In ten separate runs, all Yard. bovis isolates were thawed and cultured in 10 mL modified PPLO goop (pH seven.8) (Difco™, BD Diagnostic Systems, Sparks, Doc.), supplemented with 25% inactivated horse serum (Gibco™), 0.7% technical yeast extract, 0.5% sodium pyruvate (ReagentPlus, Sigma-Aldrich®), 0.5% D-( +)-glucose monohydrate (Sigma-Aldrich) and 0.005% phenol red. Afterward 4 days of incubation (37ºC, v% CO2) a bacterial suspension of approximately 108 CFU/mL was obtained. Bacterial Dna was obtained using the ZymoBIOMICS DNA Miniprep kit (Zymo Research) co-ordinate to the manufacturer's instructions. Quantity and quality were verified using NanoDrop ND-1000 spectrophotometer (Thermo Scientific). Low quality samples were further cleaned using CleanNGS (CleanNa) beads. All runs included the M. bovis PG45 type strain (ATCC 25523) and modified PPLO goop as the positive and negative control, respectively.
Library grooming and MinION long-read sequencing
Quality-checked native M. bovis DNA was immediately used for library preparation using the Rapid Barcoding Sequencing Kit (SQK-RBK004; Oxford Nanopore Technologies (ONT)), post-obit manufacturers' instructions. For each run, ten field strains, 1 positive control (PG45) and 1 negative control (sterile goop) were multiplexed (400 ng Deoxyribonucleic acid per sample). A new R9.iv.ane Flow cell (ONT) was used for a 48 h sequencing run on MinION device (ONT). Raw fast5 read files were nerveless using MinKnow v.3.vi.five.
Bioinformatics pipeline
All data were analyzed on an Ubuntu 18.04.three LTS system. In gild to speed up bioinformatics analyses, GPU resources (GeForce RTX 2080 Ti/PCIe/SSE2) were exploited where possible. Raw fast5 files were basecalled using Guppy basecaller (GPU v.3.3.0; ONT), followed by demultiplexing, adapter trimming, and quality filtering (Q-score ≥ 7) of fastq files with qcat (v.ane.1.0; ONT) and NanoFilt (v. ii.5.0; [30]), respectively. Reference-based assemblies were generated using the 1000. bovis PG45 blazon strain sequence (NC_014760.1) by mapping filtered reads onto the reference using GraphMap (v.0.five.2; [31]). Concluding consensus sequences were generated using Medaka (GPU v.0.10.0; ONT). All strains were identified as M. bovis using Kraken2 (v2.0.8; [32]) by adjustment the reads against the minikraken_v1_8GB database with standard settings. Overall consensus associates accuracies were verified by comparing total Single Nucleotide Polymorphisms (SNPs) using the CSI phylogeny package (v1.4, Center for Genomic Epidemiology, Denmark; [33]) as compared to the M. bovis PG45 type strain (NC_014760.1) reference sequences. To validate the use of long-read sequencing, SNPs of ten contained M. bovis PG45 assemblies were compared to those in a single MiSeq experimental dataset. All Chiliad. bovis consensus genomes are available for download on the NCBI GenBank database under the BioProject PRJNA639688 and accession numbers (SAMN15246515-SAMN1524662). Sequencing summaries can be establish in Additional file 1.
Phylogenetic analysis
Phylogenetic assay was performed on all newly generated consensus sequences solitary or in combination with 250 previously published M. bovis sequences using the FastTree-based CSI Phylogeny v1.4 (see Additional file two). All analyses included the M. bovis PG45 type strain (NC_014760.1) equally reference and 1000. agalactiae PG2 (NC_009497.1) equally outgroup. Resulting Newick files were visualized with MEGA-X software [34].
Cluster and strain determination
Due to the lack of relatedness criteria for SNP typing schemes of M. bovis and the need to establish these per organism and experimental design [35], clusters were defined by visual inspection of the phylogenetic tree and by taking into business relationship bootstrap support. In addition, the matrix of pairwise SNP counts was extracted from CSI Phylogeny for further inspection. Mean SNP differences were calculated between inside-cluster isolates, and outliers were defined with the 1.5xIQR rule, using the Outlier Reckoner (https://miniwebtool.com/outlier-calculator/).
Geographical distribution
Esri®ArcMap™ (version 10.7.1) software was used to visualize the geographical distribution and density of M. bovis isolates over Belgium. Herd size was based on the national Identification and Registration system, containing, on the first of January 2017, a total of 23 995 cattle herds in Belgium (23 733 conventional herds; 262 veal), and a full of 2 517 850 cattle. The spatial distribution of the Belgian cattle (both cattle and veal calves) was displayed using kernel smoothing. Coordinates of Belgian cattle herds were converted into a continuous raster using the kernel density estimation role weighted past number of cattle (Spatial Analyst, ArcMAP 10, ESRI, Redlands, CA, The states).
Results
Phylogenetic analysis of Belgian isolates
A median sequencing depth of 618X (range: 32X-2689X) was obtained from long reads with an average N50 read length of 5706 ± 1514 bp for all Belgian M. bovis isolates. First, implementation of long-read sequencing of M. bovis genomes was validated by comparing total SNPs from ten independently sequenced K. bovis PG45 sequences and a single Thou. bovis PG45 MiSeq dataset, showing 53 ± three SNPs and 27 SNPs difference, respectively, compared with the i 003 404 bp of the M. bovis PG45 reference genome (NC_014760.1). The observed average SNP difference of 0.005% for the long-read sequencing arroyo was considered acceptable to allow meaningful phylogenetic analyses. In improver, control strain M. bovis PG45 results were mutually compared over all 10 runs, showing a hateful SNP difference of 20 (range eight–30, standard difference 4.half-dozen), which was as well demonstrated an acceptable inter-experimental variation.
Taking into business relationship all Belgian M. bovis strains, and besides including the outgroup Grand. agalactiae, 51.four% of the M. bovis genome or 515 324 nucleotide positions were used for phylogeny. The minimum and maximum SNP differences amongst Belgian M. bovis isolates were 33 and 3775, respectively.
Visual inspection of the phylogenetic analysis of 100 Thousand. bovis isolates resulted in 5 clusters (I-V): iii large clusters (n ≥ 10 isolates), ii smaller clusters (due north < 10), and one distinct strain (VK30) as shown in Figure 1. Cluster I to Five contained 3, seven, 16, 33, and 40 M. bovis isolates, respectively. Inspection of pairwise SNP differences per cluster, showed more than homogeny within clusters II and III (mean ΔSNPs of 87 and 316, respectively), compared with cluster I, IV and V (mean ΔSNPs of 834, 1027, and 1435) (Table i). Hateful SNP differences among within-cluster isolates and outlier calculations showed outliers within cluster Iii (Mb222, Mb231), IV (Mb201, Mb240, Mb175, TOVK) and Five (Mb116, Mb166, VK11, VK23).
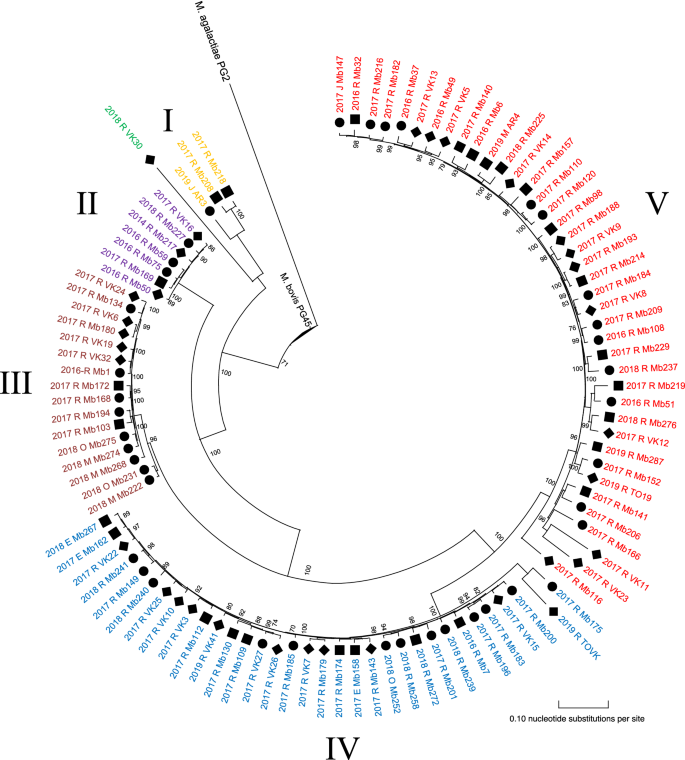
SNP-based phylogenetic tree of 100 Chiliad. bovis isolates from Belgian dairy, beef and veal farms. The figure was created using MEGA-X software with M. bovis isolates obtained over 2014–2019. The tree was rerooted to M. agalactiae PG2, which was included as an outgroup. Clusters (I-5), and VK30 are represented by different colors. The designation of the isolates features the sector (■ dairy; ● beef; ♦ veal), year of isolation (2014–2019), affected organ (R: respiratory tract; M: milk; E: ear, J: articulation and O: other) and sequence identification (run across Additional file 1). The scale bar indicates the number of substitutions per site, and bootstrap values are represented on nodes.
Betwixt and within clusters, no clan could be observed for the different cattle sectors or yr of isolation (Figure 1). Ii unlike isolates from the same herd (veal) and same sampling menstruum (Mb49 and Mb50) did not cluster together (Ii and V). All clusters persisted in Belgium for at least ane.5 sequent years throughout the country. G. bovis strains isolated from the heart ear (n = 3) were clustered within cluster Iv, while those obtained from milk, joint and other samples were scattered over different clusters (Figure 1). Finally, no clear clan between geographic location of sampled farms and Thou. bovis clusters was observed, as shown in Figure 2.
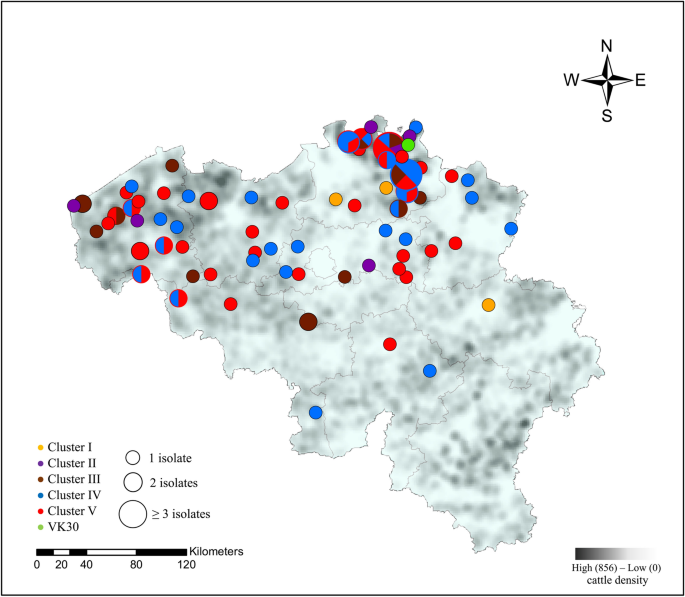
Geographical distribution of unlike G. bovis clusters over 2014–2019 and cattle density in Belgium (2017). The map was created using Esri®ArcMap™ (version ten.seven.1) software. Clusters (I-V) are represented past different colors and the radius of the circle represents the number of isolates from i hamlet. Mixed colors within one circle represent the presence of different clusters within ane hamlet.
Phylogenetic analysis of M. bovis worldwide
All 100 Belgian isolates were added to the worldwide phylogenetic tree. The percentage of the reference genome covered by all isolates, including the PG45 standard strain and the K. agalactiae outgroup strain PG2 was 39.iii%, therefore 394 303 positions were constitute in all analyzed genomes. The minimum and maximum SNP differences among all Yard. bovis isolates including the reference strain, were 0 and 4871, respectively. Belgian clusters are situated in different parts of the phylogenetic tree worldwide (Figure 3). Cluster I is related to strains isolated in the U.s.a. (2007; mean ΔSNPs of 636) and Israel (2016; mean ΔSNPs of 1369). Cluster 2 is closely related to ane recent French strain (2016; mean ΔSNPs of 171) and is situated in a larger cluster related to older strains from Israel and Eastern Europe (2001–2009; ΔSNPs < 200), and other more recently isolated strains from Israel and Eastern Europe (2011–2017; ΔSNPs < 500). Belgian cluster III and V do not cluster together with not-Belgian isolates, while cluster Four is closely related (ΔSNPs < 300 without cluster Four outliers) to M. bovis strains obtained in State of israel (2012–2017) and Eastern Europe (2013–2016). VK11 remains an outlier that does not collate with the rest of cluster V. Consequent with Figure 1, VK30 is well separated from the other Belgian isolates and is very closely related (mean ΔSNPs of 171) to strains obtained from milk in the United states of america (2017).

SNP-based topology of 350 M. bovis isolates in MEGA-X. The tree was rerooted to M. agalactiae PG2 (EPS 1952 PG2), which was included as outgroup. Belgian clusters (I-5) were collapsed every bit far as possible and represented by dissimilar colors (I: yellowish; II: purple; Three: chocolate-brown; Iv: blue, Five: cerise; VK30: light-green). The designation of the isolates contains the coded name of land of origin (ISO 3166-i; Alpha-3 code), continent of origin (● Europe; o Non-Europe), year of isolation (1952–2019), affected organ (R: respiratory tract; M: milk; E: ear, J: articulation and O: other or unknown) and sequence identification (see Boosted file 2) or the number of collapsed Belgian isolates between brackets. Bootstrap values are represented on nodes.
Discussion
In this written report, i-hundred M. bovis isolates from different Belgian cattle sectors (beef, dairy or veal) were phylogenetically compared to investigate whether sector-specific strains be and whether such strains are related to M. bovis strains previously isolated and sequenced worldwide.
In this written report, we chose to apply the ONT long-read secquencing approach [36], considering no default WGS approaches are defined for Mycoplasma spp. and short-read sequencing biases have been described for genomes with highly repetitive regions.
WGS approaches have become more than attractive over the last years, as the cost for next-generation sequencing has significantly reduced. Single Nucleotide Polymorphism (SNP) analysis using Illumina short read data from 1000. bovis isolates already showed to be an effective manner for Yard. bovis genotyping [24, 25]. Long-read nanopore sequencing (Oxford Nanopore Technologies) is known to create much faster results, and was recently practical in veterinarian medicine too [37, 38]. Nonetheless, lower unmarried read accuracies are currently obtained with ONT in comparing to Illumina [39]. Therefore, the implementation of long-read sequencing to generate M. bovis genome assemblies was verified, showing just on average 53 SNPs difference of the long-read approach with the publically available One thousand. bovis PG45 genome, representing an acceptable error rate of 0.005%. As a result, the authors believe that nanopore sequencing is a highly accessible tool, allowing applied use outside academia in routine diagnostics and real time surveillance.
From the study results, several interesting observations were fabricated. First of all, the obtained M. bovis isolates belonged to at least five different Grand. bovis clusters, of which three ascendant clusters were identified. This is in agreement with an Israeli study based on WGS-SNP, where six clusters were observed of which 3 were dominant. Remarkably, 1 cluster contained more than 50% of the isolates in that study [25]. Several other studies also showed i or two dominant lineages, although different typing methods were used [13, 15, 18, 24, 40]. In contrast to Aebi et al. [41], where mostly herd-specific K. bovis isolates were seen in Switzerland, nosotros observed close relatedness of M. bovis isolates over the different herds. This might be a outcome of more frequent purchasing cattle from unlike origins and transportation in Belgium, because 40% of cattle is transported at least once (and up to eight times) over a 5-twelvemonth lifespan in Belgium [7, 16, 17, 26, 42]. As such, the higher heterogeneity observed in cluster I, Four and Five compared with clusters II and Iii, may exist caused past different rates of genetic drift between clonal lines [17].
Secondly, no sector-specific strains or clusters were identified in the present report, which does not entirely come up as a surprise. In Belgium, veal calves are purchased from both dairy and beef farms, and fattened and slaughtered in specialized veal farms and slaughterhouses, respectively [27]. As well, approximately fifteen–20% of the farms in Belgium are mixed farms. Therefore, contact among dissimilar cattle sectors is intense. In addition, herd visitors and artificial insemination might play a role in spreading of Yard. bovis or introducing new strains on farms [43,44,45].
Thirdly, when we take a closer look at how the dissimilar clusters have spread over Belgium, no articulate association with location was observed. This was also concluded in studies performed in the UK and United states of america [17, 23, 42]. Nevertheless, in the provinces of Antwerp and Western Flanders seem to be hotspots for Grand. bovis outbreaks. As well a loftier number of local transports, Antwerp and the Flanders area are also the primary gates for cattle import, which makes these areas predisposed for the introduction of new M. bovis strains [26]. Unfortunately, M. bovis genomes were not available for isolates obtained from the top import countries for Belgium, which are Germany and kingdom of the netherlands.
Although this study was not designed to draw definitive conclusions well-nigh yr of isolation and affected organs, some preliminary observations can be made. For example, no association betwixt Grand. bovis strain and year of isolation was observed. On the other hand, nosotros saw that representatives of all clusters persisted for at least 1.5 consecutive years on Belgian territory. The persistence of strains within a land or herd has been described before [16, 24, 41]. Furthermore, shifts betwixt dominant lineages from older to new strains take been reported before as well [13, 15, 46, 47]. Also, we did non find an clan betwixt cluster and affected organ, which is in line with previous studies [14, 23, 24]. Yet, information technology was remarkable that all isolates obtained from the middle ear were clustered, which could propose the middle ear as possible predilection site for certain 1000. bovis strains. However no definitive conclusions can be drawn as there were but few isolates obtained from this isolation site in the nowadays study. In addition, nosotros isolated different Yard. bovis strains (Mb49, Mb50) from two veal calves on the same farm at the same fourth dimension. The observation of two different strains in one herd or even one animal has been described before [xiv, 16, 42, 48], in contrast to Arcangioli et al. [46], who isolated merely one identical dominant profile in the same feedlot.
Finally, it was evident that Belgian isolates were mostly related to European and Israeli M. bovis isolates, even though only a few genomes of European M. bovis isolates have been published in the NCBI database. This seems plausible, as Belgian farmers mostly purchase cattle from European farms, while State of israel too partly imports cattle from Eastern-Europe. The fact that Israeli isolates are often related to Chinese and Australian strains, is as well due to import of cattle, as outlined in detail elsewhere [25, twoscore]. Some of the Belgian outlier strains were related to American isolates, which might be explained by the fact that Yard. bovis was get-go isolated in the USA and outbreaks in Europe were only seen years afterwards. So, we can simply speculate whether these outlier strains could have been imported or evolved geographically singled-out from each other. Clusters of the Belgian isolates were not amassed exactly in the aforementioned way in the Belgian vs. the worldwide phylogenetic tree. A possible explanation could be the loss of overall coverage betwixt the construction of both phylogenetic copse (51% for the Belgian to 40% worldwide). This might be due to (ane) heterogeneity among isolates worldwide and/or (ii) the employ of genomes obtained by different laboratories, using dissimilar sequencing protocols, as the quality can exist influenced by strain maintenance, DNA extraction, library preparation, sequencing, and the bioinformatics analysis [49].
In conclusion, multiple M. bovis clusters were circulating in Belgium in 2014–2019, and were persisting for several years. Neither the veal manufacture, nor any other cattle industry could be identified as source of strain persistence. Connections between dairy, beefiness and veal industry are intense and M. bovis appears to easily spread among these sectors. The One thousand. bovis issues in the veal industry seem more than likely the consequences of strain import from dairy and beef, rather than persistence of a limited number of veal specific strains. This data can contribute to better command and prevention of Thou. bovis infections by improved biosecurity.
Availability of data and materials
All M. bovis consensus genomes are bachelor for download on the NCBI GenBank database under the BioProject PRJNA639688 and accession numbers (SAMN15246515-SAMN1524662). Sequencing summaries can be constitute in Additional file 1.
Abbreviations
- AFLP:
-
amplification fragment length polymorphism
- AMU:
-
antimicrobial apply
- AP-PCR:
-
arbitrarily primed PCR
- MLST:
-
multi-locus sequence typing
- MLVA:
-
multiple-locus variable-number tandem repeat
- ONT:
-
Oxford Nanopore Technologies
- PFGE:
-
pulsed-field gel electrophoresis
- RAPD:
-
random amplification of polymorphic Deoxyribonucleic acid
- SNPs:
-
single Nucleotide Polymorphisms
- WGS:
-
whole genome sequencing
References
-
Maunsell FP, Woolums AR, Francoz D et al (2011) Mycoplasma bovis infections in cattle. J Vet Intern Med 25:772–783. https://doi.org/10.1111/j.1939-1676.2011.0750.x
-
Maunsell FP, Donovan GA (2009) Mycoplasma bovis Infections in Immature Calves. Vet Clin North Am Food Anim Pract 25:139–177. https://doi.org/10.1016/j.cvfa.2008.10.011
-
Nicholas RAJ, Ayling RD (2003) Mycoplasma bovis: Disease, diagnosis, and control. Res Vet Sci 74:105–112. https://doi.org/x.1016/s0034-5288(02)00155-eight
-
Pull a fast one on LK, Kirk JH, Britten A (2005) Mycoplasma mastitis: a review of manual and control. J Vet Med Ser B Infect Dis Vet Public Heal 52:153–160. https://doi.org/10.1111/j.1439-0450.2005.00845.ten
-
Pardon B, De Bleecker K, Dewulf J et al (2011) Prevalence of respiratory pathogens in diseased, not-vaccinated, routinely medicated veal calves. Vet Rec. https://doi.org/ten.1136/vr.d4406
-
Pardon B (2012) Morbidity, bloodshed and drug use in white veal calves emphasis on respiratory affliction. PhD Thesis, Ghent University, Kinesthesia of Veterinary Medicine, Department of Large Animal Internal Medicine. http://hdl.handle.cyberspace/1854/LU-3007344
-
Pardon B, Callens J, Maris J et al (2020) Pathogen-specific risk factors in astute outbreaks of respiratory disease in calves. J Dairy Sci 103:2556–2566. https://doi.org/10.3168/jds.2019-17486
-
Gautier-Bouchardon AV (2018) Antimicrobial resistance in Mycoplasma spp. Microbiol Spectr half-dozen:425–446. https://doi.org/ten.1128/microbiolspec.arba-0030-2018
-
Maunsell FP, Chase C (2019) Mycoplasma bovis: Interactions with the Immune System and Failure to Generate an Effective Immune Response. Vet Clin North Am Food Anim Pract 35:471–483. https://doi.org/ten.1016/j.cvfa.2019.08.003
-
Arcangioli MA, Duet A, Meyer G et al (2008) The function of Mycoplasma bovis in bovine respiratory illness outbreaks in veal dogie feedlots. Vet J 177:89–93. https://doi.org/10.1016/j.tvjl.2007.03.008
-
Bokma J, Boone R, Deprez P, Pardon B (2019) Risk factors for antimicrobial use in veal calves and the association with mortality. J Dairy Sci 102:607–618. https://doi.org/10.3168/jds.2018-15211
-
Gautier-Bouchardon AV, Ferré S, Le Thou D et al (2014) Overall decrease in the susceptibility of Mycoplasma bovis to antimicrobials over the past xxx years in France. PLoS One 2:e87672. https://doi.org/10.1371/journal.pone.0087672
-
Becker CAM, Thibault FM, Arcangioli MA, Tardy F (2015) Loss of diversity within Mycoplasma bovis isolates collected in France from bovines with respiratory diseases over the final 35 years. Infect Genet Evol 33:118–126. https://doi.org/ten.1016/j.meegid.2015.04.019
-
Rosales RS, Churchward CP, Schnee C et al (2015) Global multilocus sequence typing assay of Mycoplasma bovis isolates reveals two chief population clusters. J Clin Microbiol 53:789–794. https://doi.org/ten.1128/JCM.01910-xiv
-
Bürki S, Spergser J, Bodmer One thousand, Pilo P (2016) A dominant lineage of Mycoplasma bovis is associated with an increased number of astringent mastitis cases in cattle. Vet Microbiol 196:63–66. https://doi.org/10.1016/j.vetmic.2016.x.016
-
Butler JA, Pinnow CC, Thomson JU et al (2001) Use of arbitrarily primed polymerase chain reaction to investigate Mycoplasma bovis outbreaks. Vet Microbiol 78:175–181. https://doi.org/10.1016/S0378-1135(00)00286-8
-
McAuliffe L, Kokotovic B, Ayling RD, Nicholas RAJ (2004) Molecular epidemiological analysis of Mycoplasma bovis isolates from the United Kingdom shows two genetically distinct clusters. J Clin Microbiol 42:4556–4565. https://doi.org/10.1128/JCM.42.10.4556-4565.2004
-
Spergser J, Macher K, Kargl Chiliad et al (2013) Emergence, re-emergence, spread and host species crossing of Mycoplasma bovis in the Austrian Alps caused by a single owned strain. Vet Microbiol 164:299–306. https://doi.org/x.1016/j.vetmic.2013.02.007
-
Maiden MCJ, Bygraves JA, Feil E et al (1998) Multilocus sequence typing: a portable approach to the identification of clones within populations of pathogenic microorganisms. Proc Natl Acad Sci U S A 95:3140–3145. https://doi.org/10.1073/pnas.95.6.3140
-
Annals KB, Lysnyansky I, Jelinski MD et al (2020) Comparison of two multilocus sequence typing schemes for Mycoplasma bovis and revision of the PubMLST reference method. J Clin Microbiol 58:e00283–e320. https://doi.org/x.1128/jcm.00283-20
-
Deng 10, den Bakker HC, Hendriksen RS (2016) Genomic epidemiology: whole-genome-sequencing–powered surveillance and outbreak investigation of foodborne bacterial pathogens. Annu Rev Food Sci Technol 7:353–374. https://doi.org/ten.1146/annurev-nutrient-041715-033259
-
Leekitcharoenphon P, Nielsen EM, Kaas RS et al (2014) Evaluation of whole genome sequencing for outbreak detection of Salmonella enterica. PLoS One 9:e87991. https://doi.org/10.1371/periodical.pone.0087991
-
Annals KB, Thole L, Rosenbush RF, Minion FC (2015) Multilocus sequence typing of Mycoplasma bovis reveals host-specific genotypes in cattle versus bison. Vet Microbiol 175:92–98. https://doi.org/10.1016/j.vetmic.2014.xi.002
-
Parker AM, Shukla A, House JK et al (2016) Genetic characterization of Australian Mycoplasma bovis isolates through whole genome sequencing analysis. Vet Microbiol 196:118–125. https://doi.org/10.1016/j.vetmic.2016.10.010
-
Yair Y, Borovok I, Mikula I et al (2020) Genomics-based epidemiology of bovine Mycoplasma bovis strains in Israel. BMC Genomics 21:seventy. https://doi.org/ten.1186/s12864-020-6460-0
-
Ensoy C, Faes C, Welby S et al (2014) Exploring cattle movements in Belgium. Prev Vet Med 116:89–101. https://doi.org/10.1016/j.prevetmed.2014.05.003
-
Pardon B, Catry B, Boone R et al (2014) Characteristics and challenges of the modernistic Belgian veal manufacture. Vlaams Diergeneeskd Tijdschr 83:155–163
-
Bokma J, Van Driessche L, Deprez P et al (2020) Rapid identification of Mycoplasma bovis from bovine bronchoalveolar lavage fluid with MALDI-TOF MS later enrichment procedure. J Clin Microbiol 58:e00004–20. https://doi.org/x.1128/jcm.00004-20
-
Bokma J, Pardon B, Van Driessche L et al (2019) Optimizing identification of Mycoplasma bovis by MALDI-TOF MS. Res Vet Sci 125:185–188. https://doi.org/x.1016/j.rvsc.2019.06.010
-
De Coster Westward, D'Hert S, Schultz DT et al (2018) NanoPack: Visualizing and processing long-read sequencing information. Bioinformatics 34:2666–2669. https://doi.org/x.1093/bioinformatics/bty149
-
Sović I, Šikić One thousand, Wilm A et al (2016) Fast and sensitive mapping of nanopore sequencing reads with GraphMap. Nat Commun vii:11307. https://doi.org/10.1038/ncomms11307
-
Woods DE, Lu J, Langmead B (2019) Improved metagenomic analysis with Kraken 2. Genome Biol twenty:257. https://doi.org/10.1186/s13059-019-1891-0
-
Kaas RS, Leekitcharoenphon P, Aarestrup FM, Lund O (2014) Solving the trouble of comparison whole bacterial genomes across different sequencing platforms. PLoS 1 ix:e104984. https://doi.org/x.1371/periodical.pone.0104984
-
Kumar S, Stecher G, Li M et al (2018) MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
-
Schürch Ac, Arredondo-Alonso Southward, Willems RJL, Goering RV (2018) Whole genome sequencing options for bacterial strain typing and epidemiologic analysis based on single nucleotide polymorphism versus factor-by-gene–based approaches. Clin Microbiol Infect 24:350–354
-
Goodwin S, McPherson JD, McCombie WR (2016) Coming of age: X years of next-generation sequencing technologies. Nat Rev Genet 17:333–351. https://doi.org/10.1038/nrg.2016.49
-
Quick J, Ashton P, Calus Due south et al (2015) Rapid draft sequencing and real-time nanopore sequencing in a hospital outbreak of Salmonella. Genome Biol 16:114. https://doi.org/10.1186/s13059-015-0677-2
-
Theuns S, Vanmechelen B, Bernaert Q et al (2018) Nanopore sequencing equally a revolutionary diagnostic tool for porcine viral enteric illness complexes identifies porcine kobuvirus as an important enteric virus. Sci Rep 8:9830. https://doi.org/x.1038/s41598-018-28180-9
-
Rang FJ, Kloosterman WP, de Ridder J (2018) From squiggle to basepair: Computational approaches for improving nanopore sequencing read accuracy. Genome Biol 19:90. https://doi.org/x.1186/s13059-018-1462-9
-
Menghwar H, He C, Zhang H et al (2017) Genotype distribution of Chinese Mycoplasma bovis isolates and their evolutionary relationship to strains from other countries. Microb Pathog 111:108–117. https://doi.org/10.1016/j.micpath.2017.08.029
-
Aebi K, Bodmer Chiliad, Frey J, Pilo P (2012) Herd-specific strains of Mycoplasma bovis in outbreaks of mycoplasmal mastitis and pneumonia. Vet Microbiol 157:363–368. https://doi.org/ten.1016/j.vetmic.2012.01.006
-
Soehnlen MK, Kariyawasam South, Lumadue JA et al (2011) Molecular epidemiological assay of Mycoplasma bovis isolates from the Pennsylvania Animal Diagnostic Laboratory showing genetic diverseness. J Dairy Sci 94:1893–1899. https://doi.org/10.3168/jds.2010-3967
-
Gonzalez RN, Sears PM, Merrill RA, Hayes GL (1992) Mastitis due to Mycoplasma in the country of New York during the period 1972–1990. Cornell Vet 82:29–forty
-
Gille L, Pilo P, Valgaeren BR et al (2016) A new predilection site of Mycoplasma bovis: Postsurgical seromas in beef cattle. Vet Microbiol 186:67–70. https://doi.org/x.1016/j.vetmic.2016.02.011
-
Haapala Five, Pohjanvirta T, Vähänikkilä N et al (2018) Semen equally a source of Mycoplasma bovis mastitis in dairy herds. Vet Microbiol 216:lx–66. https://doi.org/10.1016/j.vetmic.2018.02.005
-
Arcangioli MA, Aslan H, Tardy F et al (2012) The use of pulsed-field gel electrophoresis to investigate the epidemiology of Mycoplasma bovis in French calf feedlots. Vet J. https://doi.org/10.1016/j.tvjl.2011.05.004
-
Hata E, Harada T, Itoh Thousand (2019) Human relationship betwixt antimicrobial susceptibility and multilocus sequence type of Mycoplasma bovis isolates and development of a method for rapid detection of point mutations involved in decreased susceptibility to macrolides, lincosamides, tetracyclines. Appl Environ Microbiol 85:e00575–e619. https://doi.org/x.1128/AEM.00575-19
-
Sulyok KM, Kreizinger Z, Fekete Fifty et al (2014) Phylogeny of Mycoplasma bovis isolates from Hungary based on multi locus sequence typing and multiple-locus variable-number tandem echo analysis. BMC Vet Res 10:108. https://doi.org/ten.1186/1746-6148-10-108
-
Portmann Ac, Fournier C, Gimonet J et al (2018) A validation approach of an end-to-end whole genome sequencing workflow for source tracking of Listeria monocytogenes and Salmonella enterica. Front Microbiol ix:446. https://doi.org/x.3389/fmicb.2018.00446
Acknowledgements
We thank anybody involved in the collection of the isolates (DGZ (Animal Health Service Flanders), ARSIA, Clinic of Big Fauna Internal Medicine (Ghent University), and practicing veterinarians) and the support of the technical staff at the involved laboratories, with special thanks to Arlette Van de Kerckhove, Serge Verbanck, and Marthe Pauwels.
Funding
Jade Bokma is supported by a grant of the Belgian Federal Public Service, Health, Food Concatenation Rubber and Environment (FOD)[RF 17/6313, MALDIRESP/MA]. The use of MinION was also supported past FOD [RF17/6312, PigMinION], and the Industrial Enquiry Fund of Ghent Academy [F2018/IOF-Advanced/487]. The MALDI-TOF MS was financed by the Research Foundation Flemish region (FWO-Vlaanderen) equally Hercules project [G0H2516N, AUGE/xv/05]. The funders had no role in report design, data drove and interpretation, or the decision to submit the work for publication.
Writer data
Authors and Affiliations
Contributions
JB, NV, ST, FB and BP conceptualized the experiments and were part of the validation, formal assay, interpretation and visualization of the presented piece of work. SR assisted in the visualization of the piece of work. JB, NV, KB and JC performed the data collection. JB and NV executed the experiments. Resources were provided by HN, FH, ST, FB and BP. JB, NV, FB and BP wrote the manuscript draft, and all authors read and approved the last manuscript.
Respective writer
Ethics declarations
Ethics approval and consent to participate
Not applicable. All isolates were obtained from diagnostic samples collected by field veterinarians from clinical cases, this is in compliance with EU legislation on ethics in animal experimentation [2010/63/EU].
Consent for publication
Non applicative.
Competing interests
B.P. has received honoraria for interim as speaker or consultant for pharmaceutical (Zoetis, MSD, Vetoquinol, Dopharma, Boehringer Ingelheim, Dechra, Hipra, Ceva, Merial and Elanco) and agricultural (Boerenbond, Algoet diet) companies.
Additional data
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary data
Rights and permissions
Open Admission This article is licensed nether a Creative Commons Attribution iv.0 International License, which permits utilise, sharing, accommodation, distribution and reproduction in whatever medium or format, every bit long every bit y'all requite appropriate credit to the original author(south) and the source, provide a link to the Creative Eatables licence, and indicate if changes were made. The images or other third party material in this article are included in the commodity'due south Artistic Commons licence, unless indicated otherwise in a credit line to the material. If fabric is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted apply, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/ane.0/) applies to the information fabricated available in this commodity, unless otherwise stated in a credit line to the data.
Reprints and Permissions
About this article
Cite this article
Bokma, J., Vereecke, Northward., De Bleecker, K. et al. Phylogenomic analysis of Mycoplasma bovis from Belgian veal, dairy and beefiness herds. Vet Res 51, 121 (2020). https://doi.org/x.1186/s13567-020-00848-z
-
Received:
-
Accustomed:
-
Published:
-
DOI : https://doi.org/x.1186/s13567-020-00848-z
Keywords
- cattle
- long-read nanopore sequencing
- phylogenetic assay
- SNP analysis
- whole genome
Source: https://veterinaryresearch.biomedcentral.com/articles/10.1186/s13567-020-00848-z
0 Response to "Maping of Beef Herds in California"
Post a Comment